Atomistic and Coarse-Grained
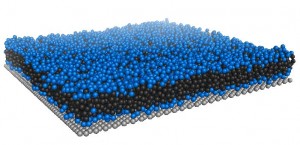
[Coarse-grained DPPC with support]
Supported lipid bilayers play a key role in understanding real biological systems but they are vastly underrepresented in computational studies.We use both coarse-grained* and atomistic simulation models to study supported and unsupported lipid bilayers. Different simplified support surfaces are prepared and placed in proximity to a lipid bilayer. The structural and dynamic properties of supported lipid bilayers are characterized. Differences between supported and unsupported bilayer systems observed in the computational study allow us to better understand the interaction between lipids and surface on a molecular level.
* J Phys Chem B 2004,108, 750-760.
Tunable Generic Coarse-Grained Model
We use a coarse-grained model** to examine the interactions between a lipid bilayer and surfaces of various shapes. In this model, a lipid consists of 3 beads and is parameterized such that no explicit water molecules are needed.
**Phys Rev E 72, 0011506 (2005)
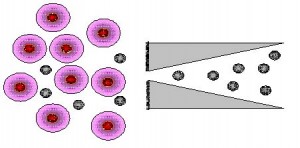
Application of Mesoscale Model
We are interested in studying the operating principles of nanoscale silicon-based biofilters in selective and gated protein transport for the purpose of protein separation. We apply a coarse grained model where proteins are treated as colloidal particles and water as a dielectric medium. Dissipative Particle Dynamics, Monte Carlo, and molecular dynamics coupled to Lattice-Boltzmann are applied to investigate the selectivity of the pore on protein transport. We investigate how the electrostatic interaction can be manipulated by the aperture size, pH, ionic strength, and applied voltage for selective passage of proteins.